By Allan Roberts
This tutorial provides instructions on downloading raw data from the Victoria Experimental Network Under the Sea (VENUS 2012) and plotting that data with the statistical application R (R Core Team 2012). For this tutorial, I will assume that you have at least some experience with R; however, if you have R installed, and can enter an instruction or two on the command line, that should be enough. If you want to plot data from the NEPTUNE Canada network, refer to my previous post, “How to plot NEPTUNE Canada data with R.” In general, downloading VENUS data is similar to downloading NEPTUNE data, but I did find that there are some differences.
Step 1. Download a data file
1.1 Go to the VENUS network website: www.venus.uvic.ca

1.2 If you already have an account click on “LOGIN.” (Note: If you have a NEPTUNE Canada account you can use the same e-mail and password to log into the VENUS site.) If you don’t have an account, click on “Register”:

1.3 Click on “DATA” and choose “DOWNLOAD DATA”:

1.4 Under “Search Type” choose “Stationary Platform”:

1.5 Under “Stationary Platform Data” choose “Search by Instrument”:

1.6 There are gaps in the data record; for this example, I’ve chosen the last two weeks of August, 2012, as an example of a particular interval with available data. Under “Time Range” set the starting and ending date and time, and then click on “Search Active Locations”:

1.7 Under “Location” choose “Saanich Inlet”, “Central Node”, and “VIP-17.” I found these to be the default settings.

1.8 Under “Instrument” choose “Aanderaa Optode 4175 (S/N 1684).” (Again, I found this to be the default setting.)

1.9 Under “Sensor(s)” choose “Temperature”:

1.10 Under “Processing” choose “No Averaging”:
1.11 Under “Data Format” choose “Comma Separated Value Text File”:

1.12 Leave “Metadata” (i.e. documentation) at the default setting of “FGDC HTML”, and click on “Search now”:

1.13 Wait ….

1.14 Click on the text file….

… you should get something like this:

1.15 Here is one place where I found that downloading VENUS data differed a bit from downloading NEPTUNE data. For the VENUS data, I’ve found that the following works: Copy and paste the downloaded data into a Word file, and then save that file as a plain text file. The plain text file is readable from R. To copy the VENUS data, click on “Select All” …

… then click on “Copy”:

1.16 Paste the file into a Word document. Click on “Save As”; make the document name “VENUS data example”; make the location your desktop, and make the format “Plain Text”:
When you save as a text file you may get the following warning. (The idea of saving as a text file is that we are not interested in fancy formatting, pictures, etc., we are only interested in the data.) Click “OK”.

The result should be a “.txt” file called “VENUS data example” on your desktop:

1.17 Create a new folder, called “VENUS network data”, and drop the text file “VENUS data example” into this folder.

Now that you have the VENUS data saved as a “.txt” file in your “VENUS network data” folder, you are ready to move onto the next step: reading the data file into R.
Step 2. Read the data file into R
The steps for changing the working directory are the same as in the NEPTUNE data tutorial:
2.1 In the R console type “getwd( )”. This will tell you what your current working directory is:

2.2 If you are on a Mac click on “Misc”, and choose “Change Working Directory”:

If you are on a PC, click on “File”, and choose “Change dir …”:

2.3 If you are on a Mac, browse for the “VENUS network data” folder, click on it, and click on “Open”; if you are on a PC browse for the “NEPTUNE data” folder, click on it, and click on “OK.”
2.4 In the R console, type “getwd( )”; the “VENUS network data” folder should now be your working directory:
2.5 Open the text file, and look at it. We want to skip all the documentation lines before the first line of data. For this example, there should be 15 lines. This includes the line with the variable names. (The variable names are not yet formatted for R; variable names in R need to be without spaces.)

2.6 Check the downloaded data by entering “head(data)”; this will show the first few lines of the data frame. (To see all the data, just type ”data”, and press enter.)

2.7 To give names to the columns, type:

Enter “head(data)” again, and you should see names at the tops of the columns. (For this tutorial, we are not concerning ourselves with the flag variable in the third column.)
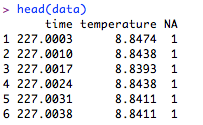
Now that the data have been downloaded, you are ready to move onto plotting …
Step 3. Plot the data
3.1 For a rough plot of the data, type:


3.2 To plot the data as a line graph with a title, and with better axis labels, we can use two steps. First: The following command will plot the data without axis labels, and make the graph a line graph in blue.

Second: Leave the graphics window open, and enter a command to add the title and axis labels.

You should get a graph like the one below. I’ve found that the easiest way to put a graph made in R into a written document or slide show is to click on the graphics window and then use copy and paste.

3.3 Now go explore the other data available from the VENUS network!
Citations
VENUS, 2012. Website. Last accessed Oct. 17, 2012: www.venus.uvic.ca
R Core Team (2012). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria.